Multimodal Data Integration: Production Architectures for Healthcare AI
- 多模态数据集成可显著提高医疗AI模型的预测能力。
- Databricks Lakehouse架构支持高效的数据管理和模型部署。
- 实际案例展示了跨数据源整合的最佳实践。
Multimodal Data Integration: Production Architectures for Healthcare AI | Databricks Blog
[](http://www.databricks.com/)
[](http://www.databricks.com/)
- Why Databricks
- * Discover
- Customers
- Partners
- Product
- * Databricks Platform
- Integrations and Data
- Pricing
- Open Source
- Solutions
- * Databricks for Industries
- Cross Industry Solutions
- Migration & Deployment
- Solution Accelerators
- Resources
- * Learning
- Events
- Blog and Podcasts
- Get Help
- Dive Deep
- About
- * Company
- Careers
- Press
- Security and Trust
- DATA + AI SUMMIT 
Table of contents
- What “governed” means in practice
- Why multimodal is becoming the default
- Four fusion strategies (and when each survives production)
- The lakehouse as a multimodal substrate
- Why the unified storage + governance model matters
- Solving the missing modality problem
- Precision oncology pattern: from architecture to clinical workflow
- Business impact: what changes when multimodal becomes operational
- Get started: a pragmatic first 30 days
Table of contents
Table of contents
- What “governed” means in practice
- Why multimodal is becoming the default
- Four fusion strategies (and when each survives production)
- The lakehouse as a multimodal substrate
- Why the unified storage + governance model matters
- Solving the missing modality problem
- Precision oncology pattern: from architecture to clinical workflow
- Business impact: what changes when multimodal becomes operational
- Get started: a pragmatic first 30 days
Healthcare & Life SciencesApril 22, 2026
Multimodal Data Integration: Production Architectures for Healthcare AI
Most multimodal healthcare AI efforts stall before production. Here’s a practical blueprint to unify genomics, imaging, clinical notes, and wearables with governance, pipelines, and fusion strategies that handle missing data.
Summary
- Build a governed multimodal foundation: Land genomics, imaging features, clinical-note entities, and wearables streams into Delta with Unity Catalog access controls, audit, lineage, and governed tags.
- Choose fusion that survives production reality: Use early/intermediate/late/attention-based fusion based on modality availability, dimensionality, and time—designed for missing modalities, not perfect cohorts.
- Operationalize end-to-end: Use Lakeflow SDP for streaming + feature windows, vector search for similarity/cohorting, and reproducible pipelines (versioning/time travel + CI/CD + MLflow) to move from POC to production.
Healthcare's most valuable AI use cases rarely live in one dataset. Multimodal data integration—combining genomics, imaging, clinical notes, and wearables—is essential for precision oncology and early detection, yet many initiatives stall before production.
Precision oncology requires understanding both molecular drivers from genomic profiling and anatomical context from imaging. Early detection improves when inherited risk signals meet longitudinal wearables. And many of the “why” details—symptoms, response, rationale—still live in clinical notes.
Despite real progress in research, many multimodal initiatives stall before production—not because modeling is impossible, but because the data and operating model aren’t ready for clinical reality. The constraint isn’t model sophistication—it’s architecture: separate stacks per modality create fragile pipelines, duplicated governance, and costly data movement that breaks down under clinical deployment needs.
This post outlines a production-oriented lakehouse pattern for multimodal precision medicine: how to land each modality into governed Delta tables, create cross-modal features, and choose fusion strategies that survive real-world missing data.
Reference architecture
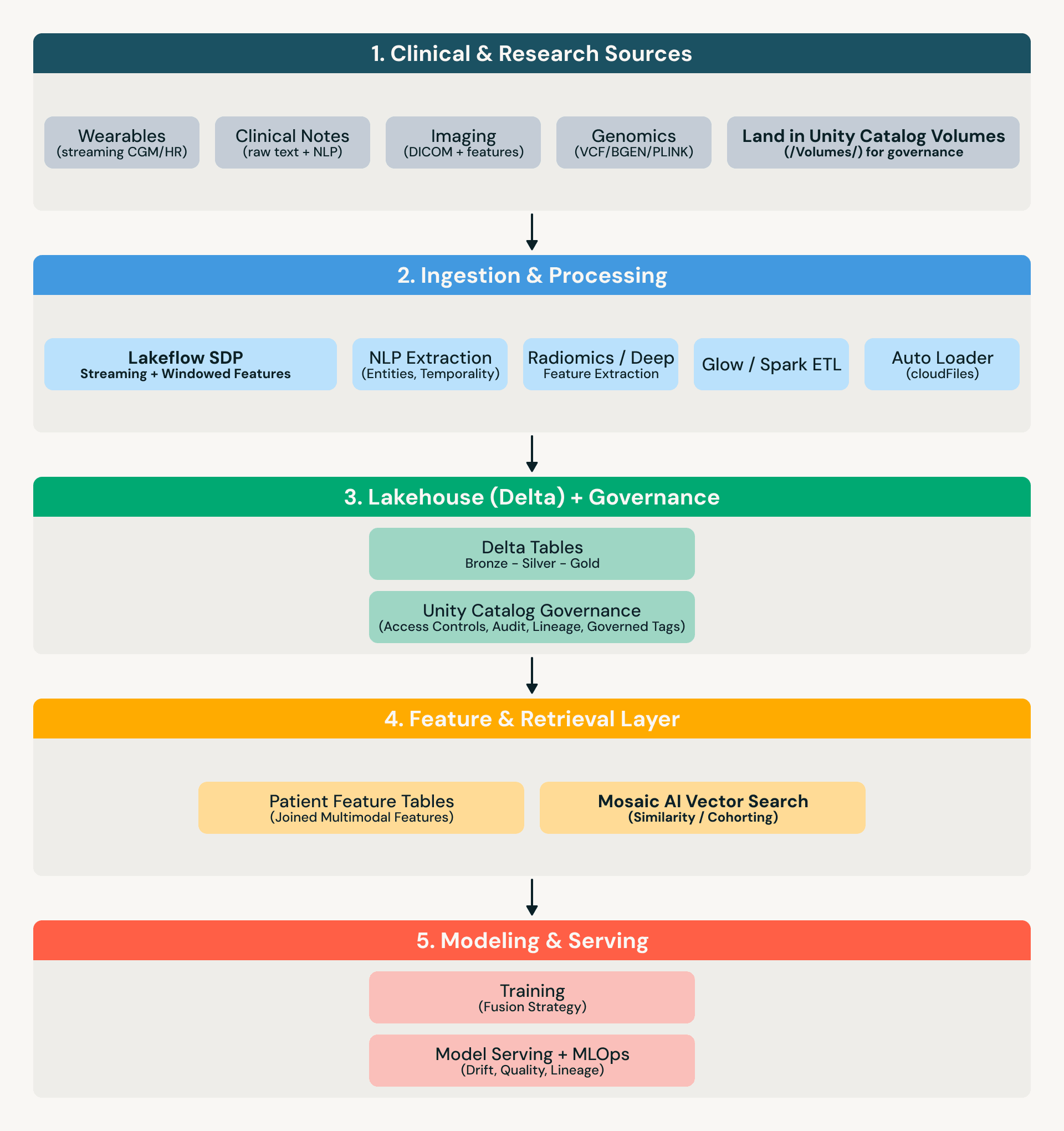
What “governed” means in practice
Throughout this post, “governed tables” means the data is secured and operationalized using Unity Catalog (or equivalent controls), including:
Data classification with governed tags: PHI/PII/28 CFR Part 202/StudyID/…
- Fine-grained access controls: catalog/schema/table/volume permissions, plus row/column-level controls where needed for PHI.
- Auditability: who accessed what, when (critical for regulated environments).
- Lineage: trace features and model inputs back to source datasets.
- Controlled sharing: consistent policy boundaries across teams and tools.
Reproducibility: versioning and time travel for datasets, CI/CD for pipelines/jobs, and MLflow for experiment and model version tracking.
This connects the technical architecture to business outcomes: fewer copies of sensitive data, reproducible analytics, and faster approvals for productionization.
Why multimodal is becoming the default
Single-modality models hit real limits in messy clinical settings. Imaging can be powerful, but many complex predictions benefit from molecular + longitudinal context. Genomics captures drivers, but not phenotype, environment, or day-to-day physiology. Notes and wearables add the “between the rows” signals that structured data often misses.
Volume reality matters: Databricks notes that roughly 80% of medical data is unstructured (for example, text and images). That’s why multimodal data integration has to handle unstructured notes and imaging at scale—not just structured EHR fields.
The practical takeaway: each modality is incomplete on its own. Multimodal systems work when they’re designed to:
1. Preserve modality-specific signal. 2. Stay robust when some inputs are missing.
Four fusion strategies (and when each survives production)
Fusion choice is rarely the only reason teams fail—but it often explains why pilots don’t translate: data is sparse, modalities arrive on different timelines, and governance requirements differ by data type.
**1) Early fusion**(Concatenate raw inputs before training.)
- Use when: small, tightly controlled cohorts with consistent modality availability.
- Tradeoff: scales poorly with high-dimensional genomics and large feature sets.
**2) Intermediate fusion**(Encode each modality separately, then merge hidden representations.)
- Use when: combining high-dimensional omics with lower-dimensional EHR/clinical features.
- Tradeoff: requires careful representation learning per modality and disciplined evaluation.
**3) Late fusion**(Train per-modality models, then combine predictions.)
- Use when: production rollouts where missing modalities are common.
- Benefit: degrades gracefully when one or more modalities are absent.
**4) Attention-based fusion**(Learn dynamic weighting across modalities and time.)
- Use when: time matters (wearables + longitudinal notes, repeated imaging) and interactions are complex.
- Tradeoff: harder to validate; requires careful controls to avoid spurious correlations.
Decision framework: match fusion to your deployment reality: modality availability patterns, dimensionality balance, and temporal dynamics.
The lakehouse as a multimodal substrate
A lakehouse approach reduces data movement across modalities: genomics tables, imaging metadata/features, text-derived entities, and streaming wearables can be governed and queried in one place—without rebuilding pipelines for each team.
Genomics processing (Glow + Delta)
Glow enables distributed genomics processing on Spark over common formats (e.g., VCF/BGEN/PLINK), with derived outputs stored as Delta tables that can be joined to clinical features.
plaintext
# Illustrative pipeline using Glow
import glow
# Ingest VCF files into Spark DataFrames
variants_df = spark.read.format("vcf").load("/Volumes/precision_med/genomics/variants/variants.vcf")
# Run GWAS at population scale (example signature; adapt to your schema)
gwas_results = glow.gwas(variants_df, phenotypes_df, covariates_df)
# Store in Delta for downstream joins with imaging + clinical features
(gwas_results.write.format("delta")
.mode("overwrite")
.saveAsTable("precision_med.genomics.gwas_results"))Imaging similarity (derived features + Vector Search)
For imaging, the pattern is: (1) derive features/embeddings upstream (radiomics or deep model outputs), (2) store features as governed Delta tables (secured via Unity Catalog), and (3) use vector search for similarity queries (e.g., “find similar phenotypes within glioblastoma”).
plaintext
from databricks.vector_search.client import VectorSearchClient
client = VectorSearchClient()
# Assumes image_embeddings were created upstream (radiomics / deep model)
client.create_index(
name="imaging_phenotypes",
source_table="precision_med.radiology.imaging_features",
primary_key="patient_id",
embedding_column="image_embeddings",
embedding_dimension=512
)
similar_cases = client.similarity_search(
query_vector=current_patient_embedding,
num_results=50,
filters={"tumor_type": "glioblastoma"}
)This enables cohort discovery and retrospective comparisons without exporting data into separate systems.
Clinical notes (NLP to governed features)
Notes often contain missing context—timelines, symptoms, response, rationale. A practical approach is to extract entities + temporality into tables (med changes, symptoms, procedures, family history, timelines), keep raw text under strict governance (Unity Catalog + access controls), and join note-derived features back to imaging and omics for modeling and cohorting.
Wearables data (Lakeflow SDP for streaming + feature windows)
**Wearables streams introduce operational requirements: schema evolution, late-arriving events, and continuous aggregation. Lakeflow Spark Declarative Pipelines (SDP) provides a robust ingestion-to-features pattern for streaming tables and materialized views. For readability, we refer to it as Lakeflow SDP below.**
Syntax note: The pyspark.pipelines module (imported as dp) with @dp.table and @dp.materialized_view decorators follows current Databricks Lakeflow SDP Python semantics.
plaintext
from pyspark import pipelines as dp
from pyspark.sql.functions import window, avg, stddev
@dp.table(
comment="Raw CGM events (streaming ingest) landed as a governed streaming table"
)
def continuous_glucose_monitor():
return (
spark.readStream
.format("cloudFiles")
.option("cloudFiles.format", "json")
.load("/Volumes/precision_med/wearables/cgm/")
.select("patient_id", "timestamp", "glucose_mg_dl", "sensor_id")
)
@dp.materialized_view(
comment="Hourly CGM feature aggregates for modeling and monitoring"
)
def glucose_variability_features():
return (
dp.read_stream("continuous_glucose_monitor")
.groupBy("patient_id", window("timestamp", "1 hour"))
.agg(
avg("glucose_mg_dl").alias("mean_glucose"),
stddev("glucose_mg_dl").alias("glucose_variability")
)
)Why the unified storage + governance model matters
The operational win is coherence:
A common failure mode in cloud deployments is a “specialty store per modality” approach (for example: a FHIR store, a separate omics store, a separate imaging store, and a separate feature or vector store). In practice, that often means duplicated governance and brittle cross-store pipelines—making lineage, reproducibility, and multimodal joins much harder to operationalize.
- Reproducibility: ACID + time travel for consistent training sets and re-analysis.
- Auditability: access logs + lineage (what data produced what feature/model).
- Security: consistent policy boundaries across modalities (PHI-safe-by-design).
- Velocity: fewer handoffs and fewer data copies across teams.
This is what turns a multimodal prototype into something you can run, monitor, and defend in production.
Solving the missing modality problem
Real deployments confront incomplete data. Not all patients receive comprehensive genomic profiling. Imaging studies may be unavailable. Wearables exist only for enrolled populations. Missingness isn’t an edge case—it’s the default.
Production designs should assume sparsity and plan for it:
- Modality masking during training: remove inputs during development to simulate deployment reality.
- Sparse attention / modality-aware models: learn to use what’s available without over-relying on any single modality.
- Transfer learning strategies: train on richer cohorts and adapt to sparse clinical populations with careful validation.
Key insight: architectures that assume complete data tend to fail in production. Architectures designed for sparsity generalize.
Precision oncology pattern: from architecture to clinical workflow
A practical precision oncology pattern looks like this:
1. Genomic profiling -> governed molecular tables (Unity Catalog). Store variants, biomarkers, and annotations as queryable tables with lineage and controlled access. 2. Imaging-derived features -> similarity + cohorting. Index imaging feature vectors for “find similar cases” and phenotype–genotype correlations. 3. Notes-derived timelines -> eligibility + context. Extract temporally-aware entities to support trial screening and consistent longitudinal understanding. 4. Tumor board support layer (human-in-the-loop). Combine multimodal evidence into a consistent review view with provenance. The goal is not to automate decisions—it’s to reduce cycle time and improve consistency in evidence gathering.
Business impact: what changes when multimodal becomes operational
Market growth is one reason this matters—but the immediate driver is operational:
- Faster cohort assembly and re-analysis when new modalities arrive.
- Fewer data copies and fewer one-off pipelines.
- Shorter iteration cycles (weeks vs. months) for translational workflows.
Patient similarity analysis can also enable practical “N-of-1” reasoning by identifying historical matches with similar multimodal profiles—especially valuable in rare disease and heterogeneous oncology populations.
Get started: a pragmatic first 30 days
1. Pick one clinical decision (e.g., trial matching, risk stratification) and define success metrics. 2. Inventory modalities + missingness (who has genomics? imaging? longitudinal wearables?). 3. Stand up governed bronze/silver/gold tables secured via Unity Catalog. 4. Choose a fusion baseline that tolerates missingness (late fusion is often a safe start). 5. Operationalize: lineage, data quality checks, drift monitoring, reproducible training sets. 6. Plan validation: evaluation cohorts, bias checks, clinician workflow checkpoints.
Keywords: multimodal AI, precision medicine, genomics processing, medical imaging AI, healthcare data integration, fusion strategies, lakehouse architecture
**High priority**
Unity Catalog: https://www.databricks.com/product/unity-catalog
Healthcare & Life Sciences: https://www.databricks.com/solutions/industries/healthcare-and-life-sciences
Data Intelligence Platform for Healthcare and Life Sciences: https://www.databricks.com/resources/guide/data-intelligence-platform-for-healthcare-and-life-sciences
**Medium priority**
Mosaic AI Vector Search Documentation: https://docs.databricks.com/en/generative-ai/vector-search.html
Delta Lake on Databricks: https://www.databricks.com/product/delta-lake-on-databricks
Data Lakehouse (glossary): https://www.databricks.com/glossary/data-lakehouse
**Additional related blogs**
Unite your Patient's Data with Multi-Modal RAG: https://www.databricks.com/blog/unite-your-patients-data-multi-modal-rag
Transforming omics data management on the Databricks Data Intelligence Platform: https://www.databricks.com/blog/transforming-omics-data-management-databricks-data-intelligence-platform
Introducing Glow (Genomics): https://www.databricks.com/blog/2019/10/18/introducing-glow-an-open-source-toolkit-for-large-scale-genomic-analysis.html
Processing DICOM images at scale with databricks.pixels: https://www.databricks.com/blog/2023/03/16/building-lakehouse-healthcare-and-life-sciences-processing-dicom-images.html
Healthcare and Life Sciences Solution Accelerators: https://www.databricks.com/solutions/accelerators
Ready to move multimodal healthcare AI from pilots to production? Explore Databricks resources for HLS architectures, governance with Unity Catalog, and end-to-end implementation patterns.
Get the latest posts in your inbox
Subscribe to our blog and get the latest posts delivered to your inbox.
Sign up
*
Work Email
*
Country Country*
By clicking “Subscribe” I understand that I will receive Databricks communications, and I agree to Databricks processing my personal data in accordance with its Privacy Policy.
Subscribe
What's next?
→
November 14, 2024/2 min read
#### Providence Health: Scaling ML/AI Projects with Databricks Mosaic AI
→
November 27, 2024/6 min read
#### How automated workflows are revolutionizing the manufacturing industry

Why Databricks
Discover
Customers
Partners
Why Databricks
Discover
Customers
Partners
Product
Databricks Platform
- Platform Overview
- Sharing
- Governance
- Artificial Intelligence
- Business Intelligence
- Database
- Data Management
- Data Warehousing
- Data Engineering
- Data Science
- Application Development
- Security
Pricing
Integrations and Data
Product
Databricks Platform
- Platform Overview
- Sharing
- Governance
- Artificial Intelligence
- Business Intelligence
- Database
- Data Management
- Data Warehousing
- Data Engineering
- Data Science
- Application Development
- Security
Pricing
Open Source
Integrations and Data
Solutions
Databricks For Industries
- Communications
- Financial Services
- Healthcare and Life Sciences
- Manufacturing
- Media and Entertainment
- Public Sector
- Retail
- View All
Cross Industry Solutions
Solutions
Databricks For Industries
- Communications
- Financial Services
- Healthcare and Life Sciences
- Manufacturing
- Media and Entertainment
- Public Sector
- Retail
- View All
Cross Industry Solutions
Data Migration
Professional Services
Solution Accelerators
Resources
Learning
Events
Blog and Podcasts
Resources
Documentation
Customer Support
Community
Learning
Events
Blog and Podcasts
About
Company
Careers
Press
About
Company
Careers
Press
Security and Trust

Databricks Inc.
160 Spear Street, 15th Floor
San Francisco, CA 94105
1-866-330-0121
- [](https://www.linkedin.com/company/databricks)
- [](https://www.facebook.com/pages/Databricks/560203607379694)
- [](https://twitter.com/databricks)
- [](https://www.databricks.com/feed)
- [](https://www.glassdoor.com/Overview/Working-at-Databricks-EI_IE954734.11,21.htm)
- [](https://www.youtube.com/@Databricks)
- [](https://www.linkedin.com/company/databricks)
- [](https://www.facebook.com/pages/Databricks/560203607379694)
- [](https://twitter.com/databricks)
- [](https://www.databricks.com/feed)
- [](https://www.glassdoor.com/Overview/Working-at-Databricks-EI_IE954734.11,21.htm)
- [](https://www.youtube.com/@Databricks)
© Databricks 2026. All rights reserved. Apache, Apache Spark, Spark, the Spark Logo, Apache Iceberg, Iceberg, and the Apache Iceberg logo are trademarks of the Apache Software Foundation.
- Privacy Notice
- |Terms of Use
- |Modern Slavery Statement
- |California Privacy
- |Your Privacy Choices
- !Image 12
We Care About Your Privacy
Databricks uses cookies and similar technologies to enhance site navigation, analyze site usage, personalize content and ads, and as further described in our Cookie Notice. To disable non-essential cookies, click “Reject All”. You can also manage your cookie settings by clicking “Manage Preferences.”
Manage Preferences
Reject All Accept All

Privacy Preference Center
Opt-Out Preference Signal Honored
Privacy Preference Center
- ### Your Privacy
- ### Strictly Necessary Cookies
- ### Performance Cookies
- ### Functional Cookies
- ### Targeting Cookies
- ### TOTHR
#### Your Privacy
When you visit any website, it may store or retrieve information on your browser, mostly in the form of cookies. This information might be about you, your preferences or your device and is mostly used to make the site work as you expect it to. The information does not usually directly identify you, but it can give you a more personalized web experience. Because we respect your right to privacy, you can choose not to allow some types of cookies. Click on the different category headings to find out more and change our default settings. However, blocking some types of cookies may impact your experience of the site and the services we are able to offer.
#### Opting out of sales, sharing, and targeted advertising
Depending on your location, you may have the right to opt out of the “sale” or “sharing” of your personal information or the processing of your personal information for purposes of online “targeted advertising.” You can opt out based on cookies and similar identifiers by disabling optional cookies here. To opt out based on other identifiers (such as your email address), submit a request in our Privacy Request Center.
#### Strictly Necessary Cookies
Always Active
These cookies are necessary for the website to function and cannot be switched off in our systems. They assist with essential site functionality such as setting your privacy preferences, logging in or filling in forms. You can set your browser to block or alert you about these cookies, but some parts of the site will no longer work.
#### Performance Cookies
- [x] Performance Cookies
These cookies allow us to count visits and traffic sources so we can measure and improve the performance of our site. They help us to know which pages are the most and least popular and see how visitors move around the site.
#### Functional Cookies
- [x] Functional Cookies
These cookies enable the website to provide enhanced functionality and personalization. They may be set by us or by third party providers whose services we have added to our pages. If you do not allow these cookies then some or all of these services may not function properly.
#### Targeting Cookies
- [x] Targeting Cookies
These cookies may be set through our site by our advertising partners. They may be used by those companies to build a profile of your interests and show you relevant advertisements on other sites. If you do not allow these cookies, you will experience less targeted advertising.
#### TOTHR
- [x] TOTHR
Cookie List
Consent Leg.Interest
- [x] checkbox label label
- [x] checkbox label label
- [x] checkbox label label
Clear
- - [x] checkbox label label
Apply Cancel
Confirm My Choices
Allow All
